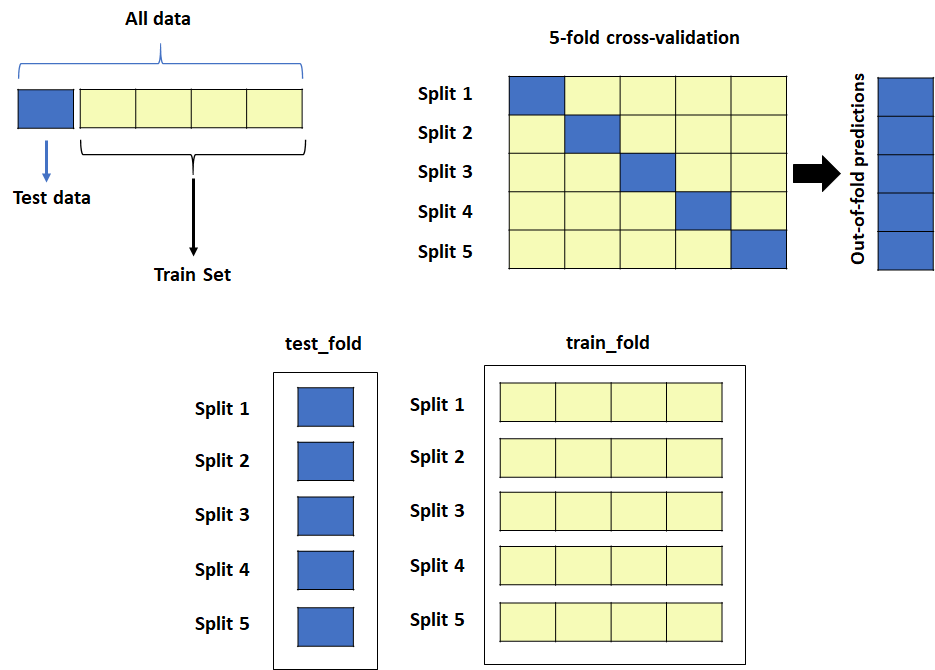
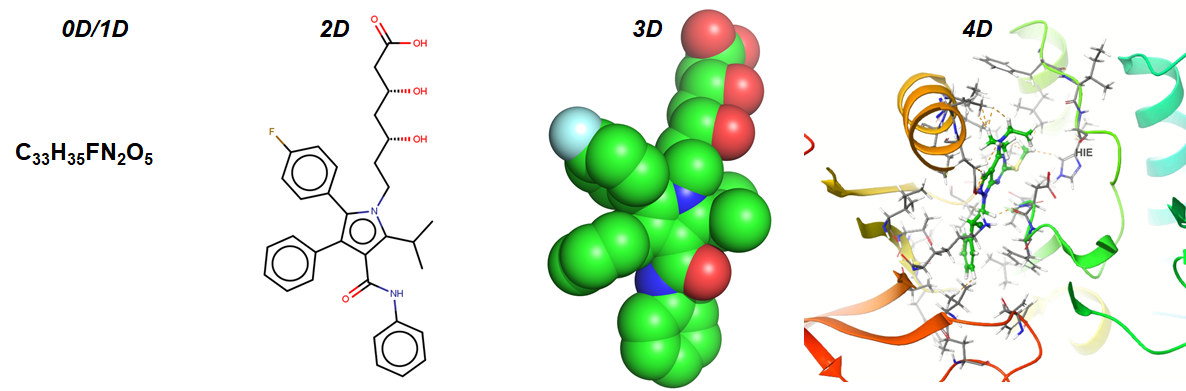
This is part 2A of the Nested Cross-Validation & Cross-Validation Series. I will go through a python tutorial on implementing k-fold CV regressors using random forest (RF) from scikit-learn with the first dataset: (A) a simple cheminformatics dataset with descriptors and endpoints of interest. In Part 2B, I will cover the same python tutorial for…